Neisseria meningitidis with decreased susceptibility to penicillin G and molecular characterization of the penA gene in strains isolated at University Hospital Centers of Casablanca and Marrakech (Morocco)
Khadija Ait Mouss, Néhémie Nzoyikorera, Aziza Razki, Bahija Zaki, Nabila Soraa, Khalid Zerouali
Corresponding author: Khadija Ait Mouss, Department of Microbiology, Laboratory of Clinical Immunology, Inflammation and Allergy, Faculty of Medicine and Pharmacy, Hassan II University of Casablanca, Casablanca, Morocco 
Received: 04 Dec 2023 - Accepted: 09 Jan 2024 - Published: 08 Feb 2024
Domain: Bacteriology,Microbiology,Molecular Biology
Keywords: Neisseria meningitidis, CMI Peni G, Sanger sequencing
©Khadija Ait Mouss et al. Pan African Medical Journal (ISSN: 1937-8688). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Khadija Ait Mouss et al. Neisseria meningitidis with decreased susceptibility to penicillin G and molecular characterization of the penA gene in strains isolated at University Hospital Centers of Casablanca and Marrakech (Morocco). Pan African Medical Journal. 2024;47:56. [doi: 10.11604/pamj.2024.47.56.42328]
Available online at: https://www.panafrican-med-journal.com//content/article/47/56/full
Research 
Neisseria meningitidis with decreased susceptibility to penicillin G and molecular characterization of the penA gene in strains isolated at University Hospital Centers of Casablanca and Marrakech (Morocco)
Neisseria meningitidis with decreased susceptibility to penicillin G and molecular characterization of the penA gene in strains isolated at University Hospital Centers of Casablanca and Marrakech (Morocco)
![]() Khadija Ait Mouss1,2,3,&,
Khadija Ait Mouss1,2,3,&, ![]() Néhémie Nzoyikorera1, Aziza Razki3, Bahija Zaki2, Nabila Soraa4,
Néhémie Nzoyikorera1, Aziza Razki3, Bahija Zaki2, Nabila Soraa4, ![]() Khalid Zerouali1,2
Khalid Zerouali1,2
&Corresponding author
Introduction: the laboratory diagnosis of meningococcal meningitis relies on conventional techniques. This study aims to evaluate the correlation between the reduced sensitivity to penicillin G of Neisseria meningitidis (N.m) strains and the expression of the altered PBP 2 gene.
Methods: out of 190 strains of N.m isolated between 2010 and 2021 at the bacteriology laboratories of Ibn Rochd University Hospital Centre (IR-UHC) in Casablanca and the UHC Mohammed VI in Marrakech, 23 isolates were part of our study. We first determined their state of sensitivity to penicillin G by E-Test strips and searched for the expression of the penA gene by PCR followed by Sanger sequencing.
Results: of all the confirmed cases of N.m, 93.15% (n=177) are of serogroup B, 75.2% (n = 143) are sensitive to penicillin G and 24.73% (n = 47) are of intermediate sensitivity. No resistance to penicillin G was observed. Reduced sensitivity to penicillin G in N.m is characterized by mutations namely F504 L, A510 V, I515 V, G541 N and I566 V located in the C-terminal region of the penA gene encoding the penicillin-binding protein 2 (PBP2) (mosaic gene).
Conclusion: our study presents useful data for the phenotypic and genotypic monitoring of resistance to penicillin G in N.m and can contribute to the analysis of genetic exchanges between different Neisseria species.
Neisseria meningitidis (N.m) commonly known as meningococcus is a strictly human pathogenic bacterium; it is presented in the form of Gram-negative diplococci. It colonizes asymptomatically the nasopharynx and can be the cause of sepsis and/or meningitis. Invasive Meningococcal Infections (IMI) require urgent diagnosis, prompt and adequate antibiotic therapy. For more than forty years [1], penicillin has been recognized as the antibiotic of choice for the treatment of these infections [2], in particular when the bacteriological diagnosis has been established [3]. Meningococcal strains resistant to penicillin G remain very rare and are associated with a production of β-lactamases, while the intermediate strains are very widely described in different countries [4]. Penicillin binding protein (PBP) are conserved proteins that play an important role in the biosynthesis of the peptidoglycan of the cell wall of many pathogenic bacteria [5,6].
Analysis by SDS-PAGE (sodium dodecyl sulfate polyacrylamide gel electrophoresis) showed that N.m contains three defined PBP genes, including PBP 1 coding for PonA, PBP 2 coding for penA and PBP 3 coding for Pbp3 [7]. The decreased sensitivity to penicillin G has been reported as being mainly due to mutations in the structure of the PBP 2 protein encoding the penA gene [8,9]. The objective of this study is to evaluate the correlation between the expression of the altered PBP 2 gene (mosaic gene) and the decreased sensitivity to penicillin G of N.m strains isolated between 2010 and 2021 in hospitalized patients.
Study design: this is a retrospective study conducted between 2010 and 2021 at the Microbiology laboratory of the IR-UHC in Casablanca and at the Bacteriology laboratory of the UHC Mohammed VI in Marrakech. These strains were analyzed in the laboratory of Invasive Meningococcal Infections of the Pasteur Institute of Morocco (IPM).
Study population: the study concerned all the services of the University Hospital Center of Casablanca and Marrakech (Morocco). Isolated samples were collected from cerebrospinal fluid (CSF) and blood cultures. Patient data consisted of age, year of isolation, ward of isolation, serogroup and MIC.
Isolate collection and identification: in total, 190 N.m isolates were collected. Twenty-three isolates with decreased sensitivity to penicillin G (0.12 ≤ MIC ≤ 0.25 µg/L) were selected for the molecular characterization of the PBP 2 gene by PCR followed by Sanger sequencing. All the strains were transplanted onto a chocolate + Polyvitex agar (bioMérieux, Marcy-l'Etoile, France). The identification was carried out according to standard bacteriological techniques.
Penicillin G sensitivity testing: the minimum inhibitory concentration (MIC) for penicillin G was determined for all isolates using the E-test strips (Oxoid Thermofisher United Kingdom) on Mueller-Hinton agar supplemented with 5% sheep's blood (bioMérieux, Marcy l'Etoile, France). The incubation was carried out at 37°C under a 5% CO2 atmosphere for 16 to 18h. The interpretation was made according to the recommendations of the European Committee on Antimicrobial Susceptibility Testing (EUCAST) depending on the year of isolation [10].
DNA extraction: DNA extraction from N.m strains was carried out by the QIAmp® DNA Minikit extraction kit (Qiagen, Hilden, Germany) according to the manufacturer's recommendations.
PCR serogroup: the serogroup of the N.m strains was confirmed by final time PCR [11,12].
PCR PenA: the reaction mixture was carried out in a final volume of 30 µl containing: MgCl 2 (5 mM); 10X PCR buffer; DNTPs (5mM); Oligonucleotides specific for penA-1F/penA-1R gene (2 mM); TaqDNA polymerase (Invitrogen Thermo Fisher Scientific) 5U/µl. The PCR search for the penA gene was carried out with the use of specific primers of a 520 bp fragment of the penA gene: penA-1F (5′GTTTTCCCAGTCACGACGTTGTAATCGAAC AGGCGACGATGTC-3′)/penA-1R (5′TTGTGAGCGGATAACAATTTCGATTAAGACG GTGTTTTGTCGG-3′). All final time PCRs were performed in the Applied Biosystems Veriti 96-well thermal cycler. The PCR amplification protocol was changed as follows: Initial denaturation at 92°C for 30 seconds followed by 20 cycles of amplification of 92°C for 30 seconds and one 30 seconds at 62°C.
PenA gene sequencing: PCR products were purified using the Ex'S-Pure kit (Image BV, Netherlands) with two hydrolytic enzymes; Exonuclease (500 µL) and Alkaline Phosphatase (SAP 500 µL) Ex'S-Pure according to the manufacturer's instructions (one cycle of 37°C for 15 min and 90°C for 10 min). Sequencing of PCR products was carried out on the ABI 3700 DNA analyzer sequencer (Applied Biosystem) with the BigDyeF® Terminator v1.1 kit (Applied Biosystems, Foster City, USA), 10X buffer, and enzymatic purifier. 25 amplification cycles are carried out on the thermal cycler; one cycle corresponds to: A DNA denaturation step at 96°C for 10 seconds. A hybridization step at 50°C for 5 seconds. A DNA elongation step at 60°C for 4 minutes [13].
Data collection: the alignment of multiple sequences of the terminal part of the penA gene and the corresponding deduced amino acid sequences was carried out by the BioEdit software (version 7.2.5). The construction and visualization of the phylogenetic tree were carried out with the programs RAxML (Randomized Axelerated Maximum Likelihood) v8.2.12 and Figtree v1.4.2 respectively. The positions of the nucleotides of the penA gene and the PBP2 amino acids are compared with the reference genome of N.m MC58 (GenBank accession number AE002098).
Ethical considerations: the isolates were obtained as part of the routine activities of the microbiology laboratory of Ibn Rochd University Hospital Center (IR- UHC) in Casablanca and the Bacteriology laboratory of the UHC Mohammed VI in Marrakech. Demographic data were extracted from the computer database anonymously. Approval of ethics and informed consent were therefore not necessary. The IMD case was defined as isolation of a N.m or meningococcal DNA detection from a normally sterile site, such as blood, CSF or articular liquid sample.
General characteristics of meningococcal isolates: of the 190 N.m strains in our study, 66.84% of the strains were isolated from cerebrospinal fluid (CSF) and 33.15% of isolates came from blood cultures. The IMIs are mainly due to serogroup B with 93.15% (n=177) of cases, followed by serogroup W with 3.70%. Only 1.57% of the isolates are from serogroup Y and 1.57% are from serogroup C. Of the strains analyzed, 143 isolates (75.26%) are sensitive to penicillin G (MIC ≤ 0.06 µg ml -1) and 24.73% (n = 47) have a reduced sensitivity (0.12 ≤ MIC ≤ 0.25 µg ml -1), most of them were from serogroup B (n=40). Of the 47 strains that phenotypically show reduced susceptibility to penicillin G, 23 (49%), show the penA gene with all five mutations, while 24 strains (51%) show no pbp2 mutation on the penA gene.
Characteristics of the penA: the amino acid sequences deduced from PBP2 (amino acids 298 to 581) were aligned in comparison with the reference strain MC58. Five mutations in 23 strains were observed at the level of pbp2 (Table 1). These modified positions were located around the preserved KTG pattern.
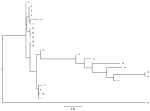
In this work, all Pen_I isolates studied, the change from amino acid Ile 566 to Val 566 (ATT to GTT) was detected at the 3' end of the PenA gene. There are four additional nucleotide transitions involving amino acid substitution, such as Phe 504 to Leu, Ala 510 to Val, Ile 515 to Val and His 541 to Asn (Figure 1). Of the set of isolates decreased susceptibility to penicillin G, 49% (n=23) of the strains analyzed had penA mosaic alleles and five amino acid substitutions (F504 L, A510 V, I515 V, G541 N and I566 V) were found in the transpeptidase region of PBP2 encoding the penA gene. Thus, 51% of the isolates sensitive to penicillin G did not carry the mosaic gene and did not contain amino acid substitutions in the PBP2 protein. The absence of the penA mosaic gene coding for the five amino acid changes associated with reduced sensitivity to penicillin G, does not necessarily indicate a penicillin G sensitive phenotype. The phylogenetic tree shows that the sequences of the penA gene are genetically diversified, and composed of four subgroups distant from the wild-type MC58 strain (Figure 2).
Invasive meningococcal disease is a worldwide public health problem, representing a major cause of morbidity and mortality. In our results, we noted that the MICs of strains with mutations coding for the five modified positions in PBP2 are increased (0.25 mg/l < MIC < 0.5 mg/l), while those strains without penA gene mutations have lower MICs.
In the mid-1980s, N.m 's resistance to antibacterial agents showed an important inflection point strains with reduced sensitivity to penicillin, which was caused by alterations in penicillin-binding protein (PBP) [14]. We know that penicillin has long been the antibiotic of choice for the treatment of meningococcal infections. The antibiotics of the β-lactam family are the most widely used, due to their good activity and with very limited side effects. In N.m, there are two mechanisms of resistance to penicillin; the first is linked to resistance by the production of β-lactamase, this mechanism characterizes some strains with an MIC greater than 1 mg/l [15]. The second resistance mechanism present in strains with a decreased sensitivity to penicillin is related to the decrease in the affinity of penicillin-binding proteins (pbp). The analysis of the sequences of the genes coding for the PBPs shows the presence of mosaic genes [15].
In the present work, 24.7% (n = 47) of the isolates are of reduced sensitivity (0.12 ≤ MIC ≤ 0.25 μg ml -1), this rate of decreased susceptibility strains is comparable to that of Portugal 24.6% [16]. The proportions of isolates exhibiting reduced sensitivity to N.m penicillin G are higher in France at 31.7% [17] and higher at 55.3% in a Spanish study [18]. In Italy, before 2002, Pen I meningococci represented 7.5% of isolates, but since then the percentage has increased to 27.4% in 2003 [19]. The strains with a decreased sensitivity to penicillin G (n=23) had penA mosaic alleles and had five amino acid substitutions (F504 L, A510 V, I515 V, G541 N and I566 V). Similar results were found in Tunisia where the majority of isolates had a reduced sensitivity to penicillin G, with a correlation between the values of resistance to penicillin and the modified penA alleles [20].
In the present work, among the 47 strains that phenotypically present a reduced sensitivity to penicillin G, 23 (49%) present the penA gene with the five mutations. Similar results were found in a large European study where the authors collected approximately 1600 isolates collected mostly from Europe, 65% showed reduced susceptibility to penicillin G, but only 38% had mosaic penA alleles [21]. In another study, on isolates of N.m with reduced penicillin susceptibility, du Plessis and his collaborators identified the mosaic penA gene in only 25 of 87 (28.7%) isolates with intermediate penicillin susceptibility [22]. In the USA, of the 466 N.m isolates collected, 10.3% (n=48) strains had reduced susceptibility to penicillin G, among of these, 63% had penA mosaic alleles, which contained all five amino acid changes [23]. In a German study, seven of 22 N. meningitidis G isolates and 19 of 75 N. lactamica strains with intermediate sensitivity to penicillin had five specific amino acid mutations in the transpeptidase domain of PBP2. The majority of intermediate strains (n = 53) of N. lactamica had only three amino acid alterations (I515 V, G541 N, and I566 V) instead of five in PBP2 [24].
In the United States between 2012 and 2016, of the 695 strains collected and characterized by whole genome sequencing, 208 isolates were of intermediate susceptibility to penicillin G, 78.8% (n=164) showed the presence of at least 4 mutations characterized in mosaic penA alleles (F504L, A510V, I515V, H541N or I566V) [25]. Reduced susceptibility to penicillin G may result from other, as yet unidentified mechanisms in addition to mutations in the penA gene. Reduced PBP-1 affinity for penicillin G and decreased expression of class 3 porin have been shown to increase penicillin G resistance in N.m [26].
The limitation of this study is related to the small sample size. The present work concerns only 23 strains with reduced sensitivity to penicillin G. The aim of this study was to evaluate the relationship between the reduced sensitivity to penicillin G of N.m strains and the expression of the altered PBP 2 gene. We suggest that future studies be conducted to evaluate other mechanisms that may reflect reduced sensitivity to penicillin G such as whole genome sequencing of N.m.
Invasive meningococcal infections are mandatory to report in Morocco; these infections require immediate diagnosis and appropriate prophylaxis. Correlation between the penA gene sequence and penicillin MIC identified mosaic structures clearly associated with reduced sensitivity. However, this is not the only mechanism that can reflect this reduced sensitivity to penicillin G but there are other mechanisms that have been described, hence the need to increasingly refer to whole genome sequencing (WGS) of N.m to understand this entire mechanism.
What is known about this topic
- Meningococcal meningitis is a serious public health problem;
- The mechanisms of penicillin resistance in N.m is linked to both resistance through β-lactamase production and decreased affinity of penicillin-binding proteins (pbp);
- Decreased susceptibility to penicillin G has been reported to be due to mutations in the structure of the pbp 2 protein encoding the penA.
What this study adds
- The frequency of N.m strains with reduced sensitivity to penicillin G has increased in recent years;
- Monitoring the relationship between penicillin MICs and penicillin sequence analysis is crucial for more effective antimicrobial and prophylactic treatment of patients and their contacts.
The authors declare no competing interests.
Conception and study design: Khalid Zerouali and Aziza Razki. Data collection: Khadija Ait Mouss, Bahija Zaki and Nabila Soraa. Data analysis and interpretation: Khadija Ait Mouss and Néhémie Nzoyikorera. Writing-review and editing: Khadija Ait Mouss, Néhémie Nzoyikorera, Aziza Razki, Bahija Zaki, Nabila Soraa and Khalid Zerouali. All the authors read and approved the final version of this manuscript.
We would like to thank all the participants of this research, Ibn Rochd University Hospital Center (IR- UHC) of Casablanca and the UHC Mohammed VI of Marrakech for their contribution to the maturation and success of this research.
Table 1: molecular characteristics of the 23 Neisseria meningitidis strains sequenced by Sanger sequencing
Figure 1: the multiple alignment of the amino acid sequences in strains with reduced sensitivity to penicillin G with that of the penA gene of the wild-type strain
Figure 2: schematic illustration of the phylogenetic analysis of the 23 Neisseria meningitidis strains isolated compared to the wild strain
- Pérez Trallero E, Garcia Arenzana JM, Ayestaran I, Muñoz Baroja I. Comparative activity in vitro of 16 antimicrobial agents against penicillin-susceptible meningococci and meningococci with diminished susceptibility to penicillin. Antimicrob Agents Chemother. 1989;33(9):1622-1623. PubMed | Google Scholar
- Oppenheim BA. Antibiotic resistance in Neisseria meningitidis. Clin Infect Dis. 1997;24 Suppl 1:S98-101. PubMed | Google Scholar
- Bertrand S, Carion F, Wintjens R, Mathys V, Vanhoof R. Evolutionary Changes in Antimicrobial Resistance of Invasive Neisseria meningitidis Isolates in Belgium from 2000 to 2010: Increasing Prevalence of Penicillin Nonsusceptibility. Antimicrob Agents Chemother. 2012;56(5):2268-2272. PubMed | Google Scholar
- Vazquez J. The resistance of Neisseria meningitidis to the antimicrobial agents: An issue still in evolution. Reviews in Medical Microbiology. 2001;12:39-45. Google Scholar
- Hill M, Deghmane A-E, Segovia M, Zarantonelli ML, Tilly G, Blancou P et al. Penicillin binding proteins as danger signals: meningococcal penicillin binding protein 2 activates dendritic cells through Toll-like receptor 4. PLoS One. 2011;6(10):e2399. PubMed | Google Scholar
- Spratt BG, Cromie KD. Penicillin-binding proteins of gram-negative bacteria. Rev Infect Dis. 1988;10(4):699-711. PubMed | Google Scholar
- Antignac A, Boneca IG, Rousselle J-C, Namane A, Carlier J-P, Vázquez JA et al. Correlation between alterations of the penicillin-binding protein 2 and modifications of the peptidoglycan structure in Neisseria meningitidis with reduced susceptibility to penicillin G. J Biol Chem. 2003;278(34):31529-31535. PubMed | Google Scholar
- Antignac A, Kriz P, Tzanakaki G, Alonso JM, Taha MK. Polymorphism of Neisseria meningitidis penA gene associated with reduced susceptibility to penicillin. J Antimicrob Chemother. 2001;47(3):285-296. PubMed | Google Scholar
- Spratt BG. Resistance to antibiotics mediated by target alterations. Science. 1994;264(5157):388-393. PubMed | Google Scholar
- Bru J-P, Caron F, Cattoen C, Cattoir V, Dubreuil L, Lina G et al. Comité de l´antibiogramme de la Société Française de Microbiologie. Recommandations 2020 V1.1. Avril. Accessed 2nd December 2023.
- Arreaza L, Vázquez JA. Molecular Approach for the Study of Penicillin Resistance In Neisseria meningitidis. Methods Mol Med. 2001;67:107-19. PubMed | Google Scholar
- Pollard AJ, Maiden MCJ. Meningococcal Disease. 2008. Springer Science & Business Media. Google Scholar
- Thulin S, Olcén P, Fredlund H, Unemo M. Total Variation in the penA Gene of Neisseria meningitidis: Correlation between Susceptibility to β-Lactam Antibiotics and penA Gene Heterogeneity. Antimicrob Agents Chemother. 2006;50(10):3317-3324. PubMed | Google Scholar
- Saez-Nieto JA, Lujan R, Berron S, Campos J, Vinas M, Fuste C et al. Epidemiology and Molecular Basis of Penicillin-Resistant Neisseria meningitidis in Spain: A 5-Year History (1985-1989). Clinical Infectious Diseases. 1992;14(2):394-402. PubMed | Google Scholar
- Bourillon A, Bingen É. Stratégie antibiotique des méningites à Neisseria meningitidis en pédiatrie. Revue Française des Laboratoires. 2004;2004(362):37-40. Google Scholar
- Ferreira E, Dias R, Caniça M. Antimicrobial susceptibility, serotype and genotype distribution of meningococci in Portugal, 2001-2002. Epidemiol Infect. 2006;134(6):1203-1207. PubMed | Google Scholar
- Antignac A, Ducos-Galand M, Guiyoule A, Pirès R, Alonso J-M, Taha M-K. Neisseria meningitidis strains isolated from invasive infections in France (1999-2002): phenotypes and antibiotic susceptibility patterns. Clin Infect Dis. 2003;37(7):912-920. PubMed | Google Scholar
- Arreaza L, de La Fuente L, Vázquez JA. Antibiotic susceptibility patterns of Neisseria meningitidis isolates from patients and asymptomatic carriers. Antimicrob Agents Chemother. 2000;44(6):1705-1707. PubMed | Google Scholar
- Stefanelli P, Fazio C, Neri A, Sofia T, Mastrantonio P. Emergence in Italy of a Neisseria meningitidis Clone with Decreased Susceptibility to Penicillin. Antimicrob Agents Chemother. 2004;48(8):3103-3106. PubMed | Google Scholar
- Brik A, Terrade A, Hong E, Deghmane A, Taha MK, Bouafsoun A et al. Phenotypic and genotypic characterization of meningococcal isolates in Tunis, Tunisia: High diversity and impact on vaccination strategies. Int J Infect Dis. 2020;91:73-78. PubMed | Google Scholar
- Taha M-K, Vázquez JA, Hong E, Bennett DE, Bertrand S, Bukovski S et al. Target Gene Sequencing To Characterize the Penicillin G Susceptibility of Neisseria meningitidis. Antimicrob Agents Chemother. 2007;51(8):2784-2792. PubMed | Google Scholar
- du Plessis M, von Gottberg A, Cohen C, de Gouveia L, Klugman KP. Neisseria meningitidis Intermediately Resistant to Penicillin and Causing Invasive Disease in South Africa in 2001 to 2005. J Clin Microbiol. 2008;46(10):3208-3214. PubMed | Google Scholar
- Harcourt BH, Anderson RD, Wu HM, Cohn AC, MacNeil JR, Taylor TH et al. Population-Based Surveillance of Neisseria meningitidis Antimicrobial Resistance in the United States. Open Forum Infect Dis. 2015;2(3):ofv117. PubMed | Google Scholar
- Karch A, Vogel U, Claus H. Role of penA polymorphisms for penicillin susceptibility in Neisseria lactamica and Neisseria meningitidis. Int J Med Microbiol. 2015;305(7):729-735. PubMed | Google Scholar
- Potts CC, Rodriguez-Rivera LD, Retchless AC, Hu F, Marjuki H, Blain AE et al. Antimicrobial Susceptibility Survey of Invasive Neisseria meningitidis , United States 2012-2016. J Infect Dis. 2022;225(11):1871-1875. PubMed | Google Scholar
- Orús P, Viñas M. Mechanisms other than penicillin-binding protein-2 alterations may contribute to moderate penicillin resistance in Neisseria meningitidis. Int J Antimicrob Agents. 2001;18(2):113-119. PubMed | Google Scholar