First report of Kingella kingae infection in a paediatric population in Accra, Ghana
Charles Addoquaye Brown, Deborah Abban, Prince Pappoe-Ashong, Alexander Martin-Odoom
Corresponding author: Alexander Martin-Odoom, School of Biomedical and Allied Health Sciences, College of Health Sciences, University of Ghana, Legon, Ghana 
Received: 26 Apr 2021 - Accepted: 26 Jul 2021 - Published: 02 Feb 2022
Domain: Bacteriology,Molecular Biology
Keywords: Kingella kingae, children, Ghana, bacteremia, blood culture, polymerase chain reaction
©Charles Addoquaye Brown et al. Pan African Medical Journal (ISSN: 1937-8688). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Charles Addoquaye Brown et al. First report of Kingella kingae infection in a paediatric population in Accra, Ghana. Pan African Medical Journal. 2022;41:95. [doi: 10.11604/pamj.2022.41.95.29528]
Available online at: https://www.panafrican-med-journal.com//content/article/41/95/full
First report of Kingella kingae infection in a paediatric population in Accra, Ghana
![]() Charles Addoquaye Brown1, Deborah Abban1, Prince Pappoe-Ashong1, Alexander Martin-Odoom1,&
Charles Addoquaye Brown1, Deborah Abban1, Prince Pappoe-Ashong1, Alexander Martin-Odoom1,&
&Corresponding author
Introduction: Kingella kingae is recognized as a frequent source of childhood bacteremia and the commonest agent of skeletal system infections in children 6 months - 4 years old. Several factors, including difficulty in detecting this fastidious organism in routine laboratory assays, result in underdiagnosis of the infections. Species-specific nucleic acid amplification assays, however, significantly improve the detection of K. kingae in blood samples. The aim of this study was to detect K. kingae infection in young children in Accra, Ghana.
Methods: a cross-sectional based study was carried out in three hospitals in Accra. Children with febrile illness and directed by a clinician for blood culture were recruited. Blood samples collected were analysed by culture and polymerase chain reaction (PCR), using universal prokaryotic and K. kingae rtxA primers.
Results: blood samples from 232 children (mean age 20.10 ± 12.57 months) were analysed. Bacteremia (72.4%) was the highest clinical diagnosis particularly in the 12-24 months age group. Only 7 (3.1%) samples showed bacterial growth and were negative for Kingella. PCR with universal prokaryotic primers succeeded in 223 (96.1%) out of 232 samples. PCR with K. kingae rtxA toxin primers was positive for 12 (5.4%) samples, all diagnosed as bacteremia, out of the 223 samples. Eleven (91.7%) out of the 12 K. kingae PCR positives were culture-negative.
Conclusion: Kingella kingae was detected only by PCR specific for the K. kingae rtxA toxin. Kingella kingae may be a potential cause of bacteremia and hence febrile illness in young children living in Accra, Ghana.
Kingella kingae is a fastidious Gram-negative coccobacillus and a normal flora of the oropharynx in young children, with a prevalence rate of 10-12% among 2 - 24-month-old children [1]. Kingella kingae is recognized as a pathogen responsible for musculoskeletal infections in the paediatric population [2, 3] and responsible for reported outbreaks of invasive K. kingae infections in the United States, Israel, and France in toddlers attending child care centres [4-8]. It is also a cause of bacterial endocarditis in children and adults [9]. Most patients with invasive K. kingae disease have moderate fever, whiles some are afebrile. Constitutional symptoms are also typically lacking, except in patients with endocarditis [9].
Detection of this fastidious organism strongly depends on the microbiologic methodology used. Kingella kingae is difficult to detect in routine laboratory assays because the fastidious nature of the bacterium makes it difficult to grow on solid medium. Also, recovery of K. kingae by culture remains unsatisfactory because K. kingae from specimen seeded onto solid culture media often results in a lack of growth [10] and thus many illnesses caused by this bacterium are probably overlooked. Results of cultures of bone aspirates and synovial fluid are often negative [11], and even if a bacterium is isolated, complete identification and antibiotic susceptibility testing may require 2 - 3 extra days. Furthermore, clinical presentation of disease due to K. kingae is often mild, thus the illness goes under diagnosed which contributes to a delay in diagnosis. This implies that, making K. kingae diagnosis requires a high index of suspicion [9].
Improvements in culture techniques and molecular detection methods have led to findings indicating that, particularly in infants and young children, K. kingae is a significantly more important pathogen than previously thought [3]. Detection of the organism approaches 100% when polymerase chain reaction (PCR) techniques are used [10]. In a comparative study by Rosey, Abachin (12), inoculation of blood culture vials and seeding samples of synovial fluid onto sheep blood agar and chocolate agar resulted in the isolation of K. kingae in 6 out of 94 (6.4%) children, whereas a combination of conventional and real-time PCR with broad-spectrum primers disclosed 15 additional cases. These results clearly demonstrated the superiority of the molecular methods and indicated that a substantial proportion of culture-negative paediatric arthritis may be attributed to K. kingae (Rosey et al., 2007). Additional studies published over the previous decade have proven that PCR enhances detection of K. kingae in samples compared with routine cultures and blood culture vials and indicated that a substantial proportion of culture-negative paediatric arthritis may be attributed to K. kingae [2, 13].
Although K. kingae is the leading agent of bone and joint infections in early childhood, the vast majority of publications on K. kingae infections have originated in countries in the developed world; reports from the developing world are still scarce. Most of the existing cases of K. kingae infection and carriage have been studied in Europe, North and South America, Australia [14], Israel [4, 15], New Zealand [16], Japan [17] and, until recently [18], none from Africa. There is not yet widespread knowledge among the medical community of K. kingae in Ghana. However, because of the potential risk for severe complications and long-term functional disability of septic arthritis and osteomyelitis in childhood, prompt laboratory confirmation and early administration of effective antimicrobial therapy are very critical to prevent late sequelae following bone and joint infections and even death. The aim of this study was therefore to detect K. kingae infection in young children in Accra, Ghana. This study provides baseline information and aids in creating the awareness of K. kingae in Ghana.
Study design: a cross-sectional study design was used.
Study sites: the study was conducted at the Greater Accra Regional Hospital (GARH), Trust Mother and Child Hospital (Trust Hospital) and Princess Marie Louise Hospital (PML), all in Accra. The PML is a quasi-private children hospital with a high attendance of children from the catchment area of Accra. The Trust Mother and Child Hospital, a satellite of the Trust Hospital, also has high attendance of children. GARH receives a lot of referral cases for children from other hospitals and clinics in Accra.
Study subjects: children 6 months to 4 years with a febrile illness or suspected localized bacterial infections (such as arthritis (joint pain) and osteomyelitis), limp, or restricted limb movement, with no history of trauma, directed by the clinician to undergo routine blood culture tests, were recruited. Children 4 years and below who reported with a febrile illness but who would not undergo routine blood culture tests were excluded.
Ethical considerations: this study was approved by the Ethics and Protocol Review Committee of the School of Biomedical and Allied Health Sciences, College of Health Sciences, University of Ghana, Korle-bu, Accra. Approval was also sought from the management of the Trust Hospital. For PML and GARH, approval was sought from the Regional Health Directorate of Greater Accra Region. Parents and/or guardians of all the children completed written informed consent forms.
Sample collection: between 1-3 ml of blood was taken from the antecubital vein of each child through venepuncture and dispensed into paediatric blood culture vials and universal blood culture bottles with brain heart infusion medium (BHI), using standard aseptic techniques [19, 20]. A drop of each blood sample (40 µl) was spotted on Whatman TM FTA cards (Whatman TM Inc., Brentford, UK) and air-dried. The Whatman TM FTA spotted cards were then kept in separate clean zipper bags for PCR analysis. The sample collection was done over five months, from April to August 2016.
Culturing
BACTEC system
The blood culture vials were sent to the Clinical Microbiology Department of the Trust Hospital. The bottles were entered into the BACTEC™ FX40 system (Becton Dickinson, Cockeysville, MD, USA) for incubation at 37°C. The BACTEC™ FX40 system reports positive blood cultures (cultures with bacteria present, thus indicating the patient is "bacteremic"). Cultures were monitored for five days, after which negative vials were removed. Positive vials were retrieved and Gram stained for a rapid general morphologic identification of the bacteria. The blood was then sub-cultured onto blood agar, MacConkey agar and chocolate agar plates. The plates were incubated at 37�C for 24 hours and isolates subjected to susceptibility testing and biochemical tests. The plates were examined for �-haemolysis, corrosion marks on the agar surface and failure to grow on MacConkey medium. For Gram-negative isolates, biochemical tests were carried out and examined, looking out for non-motility, weak oxidase reaction, a negative catalase reaction, urease and indole tests reactions, and with rare exceptions, production of acid from glucose and maltose [21].
Brain heart infusion (BHI) broth
The blood samples dispensed into BHI broth were incubated at 37°C for 24 hours. After overnight incubation, the samples were sub-cultured onto blood agar, MacConkey agar and chocolate agar plates and incubated again at 37°C in the incubator for 24 hours for bacterial growth. Gram stain was also performed on the isolates from the agar plates. Biochemical tests were performed on Gram-negative isolates. The samples were incubated again to enable slow growing bacteria to grow. After five days of incubation, the BHI sample was again sub-cultured, Gram staining was performed for a presumptive general morphologic identification of the bacteria. The plates were examined as done for the BACTEC cultures.
PCR detection of K. kingae
DNA extraction
Dried blood spots on the filter paper were cut into small pieces with scissors and transferred into 1.5-ml microtubes. For lysis, a modified salting out DNA extraction protocol with several modifications based on Asadzaheh, Javanmard (22) was used. The extraction was carried out by using TNES buffer (10 mM Tris-HCl (pH 7.5), 400 mM NaCl, 100 Mm EDTA, 0.60% SDS) protocol. For lysis, 200 �l TNES digestion buffer was added to the filter papers, followed by adding 6 �l of proteinase K (10 mg/ml) to each tube and incubation for 4 hours in a heat block (Thermo Block TDB-120, Warren, United States of America) at 56°C. Samples were retrieved, 100 μl 5M NaCl was added to each tube and mixed briefly. The contents were placed in a freezer for 5 minutes, 20 μl protein precipitator was added to each tube, then the tubes were placed in a freezer for 10 minutes and spun in a centrifuge at 15000 rpm for 10 minutes. The supernatants were transferred into new 1.5 ml Eppendorf tubes, and 400 �l 100% ethanol added to each tube, then rocked gently back and forth. Deoxyribonucleic acid (DNA) precipitation was visible at this moment. Samples were stored in a -21°C freezer for 30 minutes, retrieved, allowed to thaw, and spun down at 15000 rpm for 10 minutes. The absolute ethanol was carefully poured off the pellet, 200 �l 70% ethanol added and spun again at, 15000 rpm for 5 minutes. The 70% ethanol was poured off after spinning, and tubes were blotted and air-dried. The formed pellets were finally re-suspended in 40 μl TE buffer (10 mM Tris-HCl, pH 8.0; 1 mM EDTA, pH 8.0.) and stored at -21°C.
PCR analysis
Extracted DNA was initially subjected to PCR using universal prokaryotic primers p91E [5´-GGAATTCAAA (G/T) GAATTGACGGGGGC-3´] and p13B [5´-CGGGATCCCAGGCCCGGGAACGTATTCAC-3´] [23] to amplify a 475-bp fragment of the 16S rDNA gene. PCR was performed using OneTaq� Quick-Load� 2X Master Mix with standard buffer (NEB) and 0.25 uM of each primer. Conditions for amplification were an initial denaturation at 94°C for 4 minutes, followed by 40 cycles of 94°C for 1 minute, 56°C for 1 minute and 68°C for 2 minutes, with a final extension at 68°C for 10 minutes. Samples positive for 16S bacterial sequences were selected and used for another round of PCR using rtxA toxin gene primers of K. kingae as described previously [24]. Amplification was carried out with the primers F2-seq-rtxC [5´-GCCGAATGGGAAGATTTCTG-3´] and R2-seq-rtxA [5´-GCATTCATAAACGCCAACG-3´]. Conditions for amplification were an initial denaturation at 94°C for 4 minutes, followed by 40 cycles of 94°C for 1 minute, 56°C for 30 seconds and 68°C for 1 minute, with a final extension at 68°C for 10 minutes. All PCR reactions were carried out in a SEEAMP� SCE1000 thermal cycler (Seegene Inc., Seoul, Korea). All amplification products were visualized under short wavelength U.V. after migration in 2.0% ethidium bromide-stained agarose gels and photographed using a Kodak EDAS 290 (New York, USA) gel documentation system. The sizes of the PCR products were estimated by comparing with the mobility of a standard 100 bp DNA ladder (New England Biolabs Inc., Ipswich, MA, USA).
Data analysis
Data collected was entered and analysed using SPSS version 24.0 (IBM Corp, N.Y, USA) to obtain descriptive data. Relevant tables and figures were created from the data to allow for easy analysis and interpretation. Frequency and percentage were used to determine the prevalence of K. kingae among the children. Continuous variables such as age were presented in ranges.
Demographic data: a total of 232 children, 114 (49.1%) males and 118 (50.9%) females, were recruited for the study. Their ages ranged from 6 months to 48 months with a mean age of 20.10 ± 12.57 months.
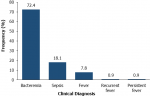
Clinical diagnosis by clinicians: the clinical diagnosis of the children by the clinicians is shown in Figure 1. Bacteremia (72.4%) was the highest diagnosis. Recurrent fever and persistent fever were the least (~1% each) diagnosis. Clinical diagnosis for any condition was highest in the 12-24 months age group (Table 1).
Culture tests: only seven (3.0%) out of the 232 culture tests were positive, showing bacterial growth. The rest showed no bacterial growth and 2 (0.9%) no bacterial growth but had contamination.
Culture results in relation to clinical diagnosis: Table 2 shows culture results in relation to clinical diagnosis. Out of the 232 samples, 168 (72.4%) were diagnosed as bacteremia of which 162 (96.4%) were culture-negative, 4 (2.4%) were culture positive and 2 (1.2%) were contamination. Of the 42 diagnosed as sepsis cases, 3 (7.14%) were positives for culture. The diagnosis for recurrent fever and persistent fever showed no bacterial growth.
PCR results
Detection of bacteria using universal prokaryotic primers: bacterial 16S rDNA fragments of the predicted sizes (Figure 2) were successfully amplified in 223 (96.1%) out of 232 samples using the universal prokaryotic p91E and p13B primers.
Detection of K. kingae using rtxA toxin gene primers: twelve (5.4%) samples yielded DNA fragments of the predicted size (approximately 1198 bp) (Figure 3) after screening of the 223 PCR positive samples for the 16S rDNA primers for K. kingae using the rtxA toxin gene primers. The 12 K. kingae PCR positives were detected in the children across all the age groups though more positives were within 12 -24 months and 25-36 months (Table 3) age groups.
Bacteria rDNA PCR in relation to culture results: out of the 223 positive 16S rDNA samples, 214 (96.0%) were culture-negative, 7 (3.1%) were positive for culture and 2 (0.9%) showed contamination.
Culture results in relation to K. kingae PCR results: comparison of culture results to K. kingae PCR results showed that out of the 12 positive K. kingae detected by PCR, 1 (8.3%) sample was culture positive and 11 (91.7%) samples were negative by the culture methods.
Clinical diagnosis in relation K. kingae PCR: Table 4 shows the comparison between the clinical diagnosis and the K. kingae PCR results. The 12 K. kingae PCR positive samples were all bacteremia cases.
In this study, K. kingae detection by culture, broad range PCR and the toxin specific rtxA genes were invesigated. Kingella kingae is a common etiology of paediatric bacteremia [2, 10]. In this study, bacteremia (72.4%) was the highest diagnosis by clinicians. However, blood culture showed bacterial growth in 7 (3.1%) samples, whereas the universal 16S rDNA detected 223 (96.1%) samples containing bacterial DNA. Detection of bacteria in a patient´s blood has diagnostic and prognostic importance, and blood cultures are essential in the diagnosis and treatment of the etiologic agents of bloodstream infections. Culturing of bacteria is the gold standard for the detection of bloodstream pathogens. Although it allows bacteria to be identified and their susceptibility profiles to be tested, it presents several limitations such as not being rapid because detection of bacterial growth requires approximately 12 - 48 h or more with fastidious bacterial [25]. Culturing also has low sensitivity for previous antibiotic treatment and/or low bacterial concentrations, due to the smaller blood sampled from paediatric patients than from adults [26, 27]. Kingella kingae may also appear Gram-positive on staining (Brujin et al., 2000: Murray et al., 2003). In addition, blood culture may allow the growth of a small quantity of bacteria potentially considered contaminants. However, many of these issues can be overcome by using nucleic acid amplification assays (NAAAs) like PCR which also reduces the time needed for bacterial identification from 3-4 days to a few hours. Using NAAAS also enables exact identification without suffering the bacteriostatic effect of antimicrobial therapy (Rothman et al., 2002: Ceroni et al., 2012). Some studies have reported detecting K. kingae from culture-negative specimens by using broad range PCR amplification (Moumile et al., 2003; Verdier et al., 2005; Matta et al., 2007; Chometon et al., 2007; Rosey et al., 2007). For example, a comparative study by Rosey et al. (2007) detected 15 additional cases using a combination of conventional and real-time PCR with broad-spectrum primers as against inoculated blood culture vials and seeded samples of synovial fluid onto solid medium which resulted in the isolation of K. kingae in 6 of 94 (6.4%) children. Using primers targeting the conserved 16S rDNA of bacteria serves as a control to monitor the extraction and absence of PCR inhibitors. PCR-based assays that amplify the 16S rRNA gene results in a 200% improvement in the diagnosis of the organism compared to culture [3]. However, several advantages of the PCR-based detection have a potential limitation of the detection of DNA from dead microorganisms, resulting in clinically false-positive results. These tests rely on all bacteria' having these 16S rDNA genes, though different bacterial species possess different numbers of copies, tied to their rate of growth [28].
Disease caused by K. kingae, believed to begin with asymptomatic colonization of the respiratory tract by disrupting the oropharyngeal epithelium which facilitates its entry into the bloodstream and damage to deeper tissues,was related to the production of a potent cytotoxin (RTX) which has a disease-promoting effect [29]. The results obtained by this study revealed that, the 223 samples that underwent PCR testing for K. kingae using the set of primers specific for rtxA gene had 12 (5.4%) being K. kingae positive; 11 (91.7%) of these were culture-negative samples. A molecular study by Ceroni et al. (2010) reported that 82% of the joint or bone aspirates of children younger than 4 years with osteoarticular infection were positive for K. kingae. PCR enhances detection of K. kingae in samples compared with routine cultures and blood culture vials [10]. Reported cases in Israel indicate that 40-50% of culture-negative septic arthritis cases in children might be attributable to K. kingae [30]. These results clearly show the superiority of molecular methods in diagnosing culture-negative paediatric arthritis (Rosey et al., 2007). The 12 Kingella kingae positive samples were all bacteremia cases by clinical diagnosis. This was not surprising as K. kingae occult bacteremia is the second most common presentation of K. kingae disease in children [9, 31]. Dubnov-Raz, Ephros (9) in their study revealed 38.8 ± 0.8�C as the maximal temperature measured in children with this condition, half with a body temperature of 39°C and one-third with a leucocyte count >15,000 WBC/ml. Thus, relying on guidelines based on the height of fever and leukocyte count results for obtaining blood cultures for management of young febrile children with no apparent focus [32], may not be sensitive enough for detecting K. kingae bacteremia.
As an oropharyngeal colonizer, K. kingae is transmitted by respiratory secretions, saliva, and potentially oral contact with contaminated objects. Asymptomatic colonization of the upper respiratory tract by K. kingae is found in children who acquire the infectious agent after six months of life [10]; subsequently, the incidence of colonization decreases to 10-12% until the end of the second year before gradually declining to low levels in older children and adults. This suggests that the disappearance of vertically transmitted immunity and the greater socialisation of children aged over six months increase the risk of colonization, whereas progressive immunological maturation leads to acquiring sufficient immunity to eradicate the organism from the pharynx in older people. These agree with the results of the present study, where more of the positive K. kingae were detected at age 1-3 years. In several studies, almost 90% of the reported cases have occurred in children aged < 5 years, and 60% in those aged < 2 years [33, 34]. Data over a 23-year period from the Soroka University Medical Centre in southern Israel indicated that age distribution of affected children show that occurrence of disease below the age of 6 months is exceptional. Cases rapidly accumulate thereafter, reaching a peak in children aged 6-11 months [9].
Kingella kingae was not detected by microbiological means but was detected only by PCR assay specific for the K. kingae RTX toxin. Most PCR positives were within the 12-36 months age group and all bacteremia cases by clinical diagnosis. Hence, K. kingae may be a potential cause of bacteremia and hence febrile illness in young children living in Accra, Ghana.
What is known about this topic
- Kingella kingae is a common etiology of paediatric bacteremia;
- Data on K. kingae infections from sub-Saharan Africa are scarce.
What this study adds
- First report of K. kingae infection in Accra, Ghana;
- Kingella kingae may be a potential cause of bacteremia in young children in Accra, Ghana.
The authors declare no competing interests.
Charles Addoquaye Brown: conception, study design, analysis and interpretation of data; Deborah Abban: data collection and analysis and interpretation of data; Prince Pappoe-Ashong: data collection and analysis and interpretation of data; Alexander Martin-Odoom: analysis and interpretation of data; Charles Addoquaye Brown, Prince Pappoe-Ashong and Alexander Martin-Odoom: drafting and revising the article; Charles Addoquaye Brown and Alexander Martin-Odoom: final approval of the version to be published. All authors have contributed immensely towards the contents of the manuscript and have all read and approved the final draft.
We are grateful to laboratory staff of Greater Accra Regional Hospital, Trust Mother and Child Hospital (Trust Hospital) and Princess Marie Louise Hospital (PML).
Table 1: clinical diagnosis in relation to age group
Table 2: comparison of culture results and clinical diagnosis
Table 3: age group and K. kingae PCR results
Table 4: comparison of clinical diagnosis and K. kingae PCR results
Figure 1: clinical diagnosis of the children by the clinicians
Figure 2: ethidium bromide-stained 2.0% agarose gel electrophoregram of amplified 16S rDNA fragments (475 bp) from samples using the p91E and p13B primers; Lane M = 100 bp marker (New England Biolabs Inc., Ipswich, MA, USA). Lanes 2 and 4 - 9 = PCR positives for 16S rDNA genes; Lanes 1 and 3 = PCR negatives for16S rDNA genes
Figure 3: ethidium bromide-stained 2.0% agarose gel electrophoregram of amplified K. kingae rtxA toxin gene fragments (1198 bp) using the rtxA primers; Lane 8= 100 bp marker (New England Biolabs Inc., Ipswich, MA, USA); Lanes 1 and 2 = PCR positive for K. kingae; Lanes 3 - 7 = PCR negative for K. kingae
- Amit U, Flaishmakher S, Dagan R, Porat N, Yagupsky P. Age-dependent carriage of Kingella kingae in young children and turnover of colonizing strains. J Pediatric Infect Dis Soc. 2014 Jun;3(2):160-2. PubMed | Google Scholar
- Yagupsky P. Kingella kingae: carriage, transmission, and disease. Clin Microbiol Rev. 2015 Jan;28(1):54-79. PubMed | Google Scholar
- Yagupsky P. Diagnosing Kingella kingae infections in infants and young children. Expert Rev Anti Infect Ther. 2017 Oct;15(10):925-934. PubMed | Google Scholar
- Yagupsky P, Erlich Y, Ariela S, Trefler R, Porat N. Outbreak of Kingella kingae skeletal system infections in children in daycare. Pediatr Infect Dis J. 2006 Jun;25(6):526-32. PubMed | Google Scholar
- Kiang KM, Ogunmodede F, Juni BA, Boxrud DJ, Glennen A, Bartkus JM, et al. Outbreak of osteomyelitis/septic arthritis caused by Kingella kingae among child care center attendees. Pediatrics. 2005 Aug;116(2):e206-13. PubMed | Google Scholar
- Yagupsky P, Ben-Ami Y, Trefler R, Porat N. Outbreaks of invasive Kingella kingae infections in closed communities. J Pediatr. 2016 Feb;169:135-9.e1. PubMed | Google Scholar
- Bidet P, Tran Quang V, Yagusky P, Birgy A, Bonacorsi S, Basmaci R. Investigation of Kingella kingae invasive infection outbreaks in day care facilities: assessment of a rapid genotyping tool targeting the DNA uptake sequence. J Clin Microbiol. 2017 Aug;55(8):2422-2430. PubMed | Google Scholar
- Seña AC, Seed P, Nicholson B, Joyce M, Cunningham CK. Kingella kingae endocarditis and a cluster investigation among daycare attendees. Pediatr Infect Dis J. 2010 Jan;29(1):86-8. PubMed | Google Scholar
- Dubnov-Raz G, Ephros M, Garty B-Z, Schlesinger Y, Maayan-Metzger A, Hasson J et al. Invasive pediatric Kingella kingae Infections: a nationwide collaborative study. Pediatr Infect Dis J. 2010 Jul;29(7):639-43. PubMed | Google Scholar
- Yagupsky P, Porsch E, St Geme JW, 3rd. Kingella kingae: an emerging pathogen in young children. Pediatrics. 2011 Mar;127(3):557-65. PubMed | Google Scholar
- Hujazi I, Oni D, Arora A, Muniz G, Khanduja V. The fate of acutely inflamed joints with a negative synovial fluid culture. Int Orthop. 2012 Jul;36(7):1487-92. PubMed | Google Scholar
- Rosey A-L, Abachin E, Quesnes G, Cadilhac C, Pejin Z, Glorion C et al. Development of a broad-range 16S rDNA real-time PCR for the diagnosis of septic arthritis in children. J Microbiol Methods. 2007 Jan;68(1):88-93. PubMed | Google Scholar
- El Houmami N, Bzdrenga J, Durand GA, Minodier P, Seligmann H, Prudent E et al. Molecular Tests That Target the RTX Locus Do Not Distinguish between Kingella kingae and the Recently Described Kingella negevensis Species. J Clin Microbiol. 2017 Oct;55(10):3113-3122. PubMed | Google Scholar
- Basmaci R, Bidet P, Yagupsky P, Muñoz-Almagro C, Balashova NV, Doit C et al. Major intercontinentally distributed sequence types of Kingella kingae and development of a rapid molecular typing tool. J Clin Microbiol. 2014 Nov;52(11):3890-7. PubMed | Google Scholar
- Dubnov-Raz G, Scheuerman O, Chodick G, Finkelstein Y, Samra Z, Garty B-Z. Invasive Kingella kingae infections in children: clinical and laboratory characteristics. Pediatrics. 2008 Dec;122(6):1305-9. PubMed | Google Scholar
- Olijve L, Podmore R, Anderson T, Walls T. High rate of oropharyngeal Kingella kingae carriage in New Zealand children. J Paediatr Child Health. 2016 Dec;52(12):1081-1085. PubMed | Google Scholar
- Kuzumoto K, Kubota N, Saito Y, Fujioka F, Yumoto K, Hidaka E et al. A case of osteomyelitis due to Kingella kingae. Kansenshogaku Zasshi. 2013 Mar;87(2):207-10. PubMed | Google Scholar
- El Houmami N, Ceroni D, Codjo Seignon K, Pons J-C, Lambert C, Durand GA et al. Acute septic arthritis of the knee caused by Kingella kingae in a 5-Year-old Cameroonian boy. Front Pediatr. 2017 Nov 6;5:230. PubMed | Google Scholar
- Yodoshi T, Ueda S, Goldman RD. Skin preparation for prevention of peripheral blood culture contamination in children. Pediatr Int. 2019 Jul;61(7):647-651. PubMed | Google Scholar
- Kiyoyama T, Tokuda Y, Shiiki S, Hachiman T, Shimasaki T, Endo K. Isopropyl alcohol compared with isopropyl alcohol plus povidone-iodine as skin preparation for prevention of blood culture contamination. J Clin Microbiol. 2009 Jan;47(1):54-8. PubMed | Google Scholar
- Odum L, Frederiksen W. Identification and characterization of Kingella kingae. Acta Pathol Microbiol Scand B. 1981 Oct;89(5):311-5 P. PubMed | Google Scholar
- Asadzaheh N, Javanmard A, Nassiry MR. Comparison of rapid DNA extraction techniques for conventional PCR-RFLP analysis from mammalian whole blood cells. J Mol Genet 2010;2(3): 32-35.
- Matta M, Wermert D, Podglajen I, Sanchez O, Buu-Hoï A, Gutmann L et al. Molecular diagnosis of Kingella kingae pericarditis by amplification and sequencing of the 16S rRNA gene. J Clin Microbiol. 2007 Sep;45(9):3133-4. PubMed | Google Scholar
- Lehours P, Freydière A-M, Richer O, Burucoa C, Boisset S, Lanotte P et al. The rtxA toxin gene of Kingella kingae: a pertinent target for molecular diagnosis of osteoarticular infections. J Clin Microbiol. 2011 Apr;49(4):1245-50. PubMed | Google Scholar
- Peters RPH, van Agtmael MA, Danner SA, Savelkoul PHM, Vandenbroucke-Grauls CMJE. New developments in the diagnosis of bloodstream infections. Lancet Infect Dis. 2004 Dec;4(12):751-60. PubMed | Google Scholar
- Randolph AG, Brun-Buisson C, Goldmann D. Identification of central venous catheter-related infections in infants and children. Pediatr Crit Care Med. 2005 May;6(3 Suppl):S19-24. PubMed | Google Scholar
- Mermel LA, Maki DG. Detection of bacteremia in adults: consequences of culturing an inadequate volume of blood. Ann Intern Med. 1993 Aug 15;119(4):270-2. PubMed | Google Scholar
- Fenollar F, Lévy PY, Raoult D. Usefulness of broad-range PCR for the diagnosis of osteoarticular infections. Curr Opin Rheumatol. 2008 Jul;20(4):463-70. PubMed | Google Scholar
- Kehl-Fie TE, St Geme JW, 3rd. Identification and characterization of an RTX toxin in the emerging pathogen Kingella kingae. J Bacteriol. 2007 Jan;189(2):430-6. PubMed | Google Scholar
- Yagupsky P, Bar-Ziv Y, Howard CB, Dagan R. Epidemiology, etiology, and clinical features of septic arthritis in children younger than 24 months. Arch Pediatr Adolesc Med. 1995 May;149(5):537-40. PubMed | Google Scholar
- Yagupsky P, Dagan R. Kingella kingae bacteremia in children. Pediatr Infect Dis J. 1994 Dec;13(12):1148-9. PubMed
- Baraff LJ, Bass JW, Fleisher GR, Klein JO, McCracken GH, Jr., Powell KR et al. Practice guideline for the management of infants and children 0 to 36 months of age with fever without source. Agency for Health Care Policy and Research. Ann Emerg Med. 1993 Jul;22(7):1198-210. PubMed | Google Scholar
- Verdier I, Gayet-Ageron A, Ploton C, Taylor P, Benito Y, Freydiere AM et al. Contribution of a broad range polymerase chain reaction to the diagnosis of osteoarticular infections caused by Kingella kingae: description of twenty-four recent pediatric diagnoses. Pediatr Infect Dis J. 2005 Aug;24(8):692-6. PubMed | Google Scholar
- Yagupsky P. Kingella kingae: from medical rarity to an emerging paediatric pathogen. Lancet Infect Dis. 2004 Jun;4(6):358-67. PubMed | Google Scholar
Search
This article authors
On Pubmed
On Google Scholar
Citation [Download]
Navigate this article
Similar articles in
Key words
Tables and figures
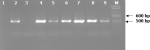 Figure 2: ethidium bromide-stained 2.0% agarose gel electrophoregram of amplified 16S rDNA fragments (475 bp) from samples using the p91E and p13B primers; Lane M = 100 bp marker (New England Biolabs Inc., Ipswich, MA, USA). Lanes 2 and 4 - 9 = PCR positives for 16S rDNA genes; Lanes 1 and 3 = PCR negatives for16S rDNA genes
Figure 2: ethidium bromide-stained 2.0% agarose gel electrophoregram of amplified 16S rDNA fragments (475 bp) from samples using the p91E and p13B primers; Lane M = 100 bp marker (New England Biolabs Inc., Ipswich, MA, USA). Lanes 2 and 4 - 9 = PCR positives for 16S rDNA genes; Lanes 1 and 3 = PCR negatives for16S rDNA genes
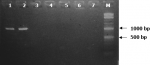 Figure 3: ethidium bromide-stained 2.0% agarose gel electrophoregram of amplified K. kingae rtxA toxin gene fragments (1198 bp) using the rtxA primers; Lane 8= 100 bp marker (New England Biolabs Inc., Ipswich, MA, USA); Lanes 1 and 2 = PCR positive for K. kingae; Lanes 3 - 7 = PCR negative for K. kingae
Figure 3: ethidium bromide-stained 2.0% agarose gel electrophoregram of amplified K. kingae rtxA toxin gene fragments (1198 bp) using the rtxA primers; Lane 8= 100 bp marker (New England Biolabs Inc., Ipswich, MA, USA); Lanes 1 and 2 = PCR positive for K. kingae; Lanes 3 - 7 = PCR negative for K. kingae