Artificial Intelligence (AI) in medicine as a strategic valuable tool
Andreas Larentzakis, Nik Lygeros
Corresponding author: Andreas Larentzakis, First Department of Propaedeutic Surgery, Athens Medical School, National and Kapodistrian University of Athens, Hippocration General Athens Hospital, Athens, Greece 
Received: 03 Feb 2021 - Accepted: 05 Feb 2021 - Published: 17 Feb 2021
Domain: Health system development,Medical informatics,Health promotion
Keywords: Artificial intelligence, healthcare, medicine, neural network, deep learning, machine learning
©Andreas Larentzakis et al. Pan African Medical Journal (ISSN: 1937-8688). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Andreas Larentzakis et al. Artificial Intelligence (AI) in medicine as a strategic valuable tool. Pan African Medical Journal. 2021;38:184. [doi: 10.11604/pamj.2021.38.184.28197]
Available online at: https://www.panafrican-med-journal.com//content/article/38/184/full
Artificial intelligence (AI) in medicine as a strategic valuable tool
&Corresponding author
Humans' creativity led to machines that outperform human capabilities in terms of workload, effectiveness, precision, endurance, strength, and repetitiveness. It has always been a vision and a way to transcend the existence and to give more sense to life, which is precious. The common denominator of all these creations was that they were meant to replace, enhance or go beyond the mechanical capabilities of the human body. The story takes another bifurcation when Alan Turing introduced the concept of a machine that could think, in 1950. Artificial intelligence, presented as a term in 1956, describes the use of computers to imitate intelligence and critical thinking comparable to humans. However, the revolution began in 1943, when artificial neural networks was an attempt to exploit the architecture of the human brain to perform tasks that conventional algorithms had little success with. Artificial intelligence is becoming a research focus and a tool of strategic value. The same observations apply in the field of healthcare, too. In this manuscript, we try to address key questions regarding artificial intelligence in medicine, such as what artificial intelligence is and how it works, what is its value in terms of application in medicine, and what are the prospects?
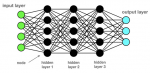
It has always been an area of challenge for humans to create machines to outperform human capabilities in terms of workload, effectiveness, precision, endurance, strength, and repetitiveness. It is a way to transcend the existence and to give more sense to life, which is precious. The common denominator of all these creations was that they were meant to replace, enhance or go beyond the mechanical capabilities of the human body. This path of evolution is smooth and predictable. This story takes a different shift or another bifurcation to be more precise when Alan Turing introduced the concept of a machine that could achieve human-level performance in thinking in 1950 [1]. However, the revolution began with the computational model for neural networks (NNs) with Warren McCulloch and Walter Pitts, and this time the evolution is unpredictable [2]. In mathematical terms, the network forms a directed, weight graph. This point of view was reinforced by Norbert Wiener, who introduced the feedback [3]. Artificial neural network (ANN) started at first level as an attempt to exploit the architecture of the human brain to perform tasks that conventional algorithms had little success with. Artificial neural network architecture is based on nodes arranged in layers and connected via their input(s) and output(s), in a way attempting to imitate brain neurons activity (Figure 1). Artificial intelligence (AI) as a term describes the use of computers to imitate intelligence and critical thinking comparable to humans, and it was first mentioned by John McCarthy during a conference held in 1956 [4].
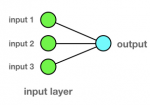
How it works: but how it works? Let us take a brain neuron; if the incoming synaptic stimuli (inputs) are of sufficient intensity, then the neuron will fire (output). Figure 2 shows a model of a single artificial neuron with three inputs and one output. Inputs and outputs are ''0'' or ''1''. In order to keep things simple, the following example will not use a firing threshold. We want to train the neuron according to the following pattern:
Case: [input a - input b - input c] ➝[output]
Case A: [0 - 0 - 0]➝[0]
Case B: [0 - 0 - 1] ➝[1]
Case C: [1 - 1 - 0] ➝[0]
Case D: [1 - 0 - 0] ➝[0]
The first step is to weight each case input by multiplying it with a random positive or negative number. Then we add all the weighted inputs of each case. Next thing, we normalize this sum of each case by using a sigmoid function in order to get a result between 0 and 1, as an output of the neuron for each case. Now we calculate the error between the normalized sum and the actual training output of each case; we use this error to adjust the weights to be used for the next round of calculations. Adjusting the weights takes under consideration the input, the calculated output, and the magnitude of the error, in a way that the adjustment to be proportional to the magnitude of the error (sigmoid curve gradient works for this). By repeating this cycle thousands of times, the neuron finally makes almost no adjustments to the weights of the inputs, meaning that it has been trained to recognize the pattern. Now one may introduce a new set of three inputs, and the already trained algorithm will be able to provide an output that corresponds to the already recognized pattern. Thus, one could summarize that artificial thinking is a pattern recognition by weighting, comparing, and adjusting many many times before a pattern replication output is created.
A sophisticated AI algorithm needs to be exposed to data feeds, which are structured and labelled in a way the algorithm can recognize (i.e. numbers, pixels, colours). Ng and Dean, Stanford and Google, respectively, leaders on computer science, created an ANN that learned to recognize higher-level concepts, such as human face, human body, or animals [5]. Unsupervised pre-training, increased computing power from multiple graphics process units (GPUs), and distributed computing allowed the use of larger (increased number of nodes) and deeper (increased number of layers) networks, particularly in image and visual recognition tasks, which became known as deep learning (DL) [6-8]. And now at high-level research, we use deep neural networks (DNNs) with tensor processing units (TPUs) [9]. Nowadays, the most representative area of thinking machines evolution has been the world of strategy board games. Board games, such as chess, shogi or go, are considered an expression of human intellect at the highest level; however, DNNs as AlphaGo, AlphaGo Master, AlphaGo Zero mastered all those sharp games [10-13]. The 3D models of proteins that AlphaFold generates are far more accurate than any that have come before marking significant progress on one of the core challenges in biology [14, 15].
Artificial intelligence applications in healthcare: machine learning (ML) algorithms based on NNs have already been used in the field of healthcare, mainly in medical diagnosis and prognosis, disease treatment, drug development, gene editing, and personalized medicine.
Disease diagnosis and prognosis: medical imaging plays a key role as an input. Plain film X-rays have been widely used as inputs in ML algorithms to teach them to diagnose lung conditions, such as pneumonia, emphysema, and tuberculosis or to detect bone age, maturity, and fractures [16-20]. Neural networks, fed with chest computed tomography (CT) scans from smokers can identify and stage chronic obstructive pulmonary disease as well as predict mortality [21]. In the field of ophthalmology, AI-based algorithms have been utilized for fundus screening in diabetic patients, age-related macular degeneration, and congenital cataract diagnosis [22-27]. Cancer diagnosis is another field that ML and NNs have been tested and proved to be superior or non-inferior to humans, including malignancy detection in pathology images, in screening mammography, in CT or magnetic resonance imaging (MRI) or positron emission tomography (PET) scans, and skin clinical images [28-40]. Also, ML algorithms fed with endoscopic images and videos reached human-like performance in gastrointestinal neoplasms detection, such as of oesophagal cancer, gastric cancer, and large bowel polyps [41-43]. Furthermore, cardiologists are investigating the ML NNs algorithms in the diagnosis, severity classification, and prognosis of cardiovascular diseases, by processing data obtained from electronic health records (EHR), electrocardiography, echocardiography, coronary artery calcium scoring, coronary CT angiography, and MRI [44-47]. For example, AI models can predict survival outcomes given a specific diagnosis, such as pulmonary hypertension by 3D cardiac MRI processing [48]. Many studies are also existing in the field of neuroscience. Deep neural networks (DNNs) can predict the future diagnosis of autism in high-risk children by processing brain magnetic resonance imaging (MRI), assess the progression of dementia by processing a single amyloid PET scan, detect intracranial haemorrhage on CTs, as well as to diagnose schizophrenia and predict the risk of suicide by the processing of functional MRIs (fMRIs) and EHR [49-54]. Finally, timely diagnosis of infectious diseases in terms of pathogen identification and antibiotic susceptibility testing is feasible through ML processing of bacterial Raman spectra or bacterial and viral mRNA [55,56].
Disease treatment: in the field of psychiatry, researchers used functional magnetic resonance imaging or functional (fMRI) and proton magnetic resonance spectroscopy (1H-MRS) as inputs to a linguistic AI platform; as a result, they were able to manage lithium dosage in bipolar patients [57]. In another study, AI virtual interviewer could capture more post-traumatic stress symptoms from veterans than the human interviewers [58]. Moreover, in the field of surgery, as surgical robots are already here, artificial intelligence short guide ribonucleic acid implementation in operations is already happening in experimental and dental settings [59,60].
Drug development: the development of a new drug is a costly and time-consuming process, which includes identification of targets for intervention, hypothesis for a new compound, and clinical trials of level I, II, and III [61]. The recognition of a possible target and the hypotheses generation for a new compound relies on pattern recognition. Chemists are skilled to recognize such patterns, relate them to retrosynthetic analysis, and predict the properties, absorption, distribution, metabolism, excretion, and toxicity (ADMET). Deep learning architecture algorithms are up-and-coming tools in the field of drug development because they imitate chemists' pattern recognition skills. Moreover, it seems possible to advance the whole process to a next level by being able to de novo design of drugs, considering all the available domain, ligand-based, and associations data during the development of a model [62,63]. The most successful paradigm of such an effort is the discovery of a new type of antibiotic, halicin, that has a different structure from known antibiotics and a broad-spectrum antibacterial activity including resistant strains such as pan-resistant Acinetobacter baumannii. The same DL algorithm was able to identify eight compounds with antibacterial activity and different structure comparing with the already known antibiotics [64]. Even if the main focus of DL-aided drug innovation is on small molecules, some approaches utilize DL to design proteins and develop antibodies [14,15,65].
Biomarkers: the principles, approaches, and tools used in drug development are applied to the identification of biomarkers, which are molecules that when found in body fluids or tissues are pathognomonic, i.e. they provide absolute certainty for disease diagnosis. Biomarkers are useful in imaging, early diagnosis, prognosis, disease progression evaluation, risk assessment for developing a specific disease, and predicting patients' response to a drug. Pembrolizumab for malignancies carrying a specific genetic biomarker is an example of how AI-aided biomarker identification could lead to the development of targeted biotherapies [66]. There are more other AI biomarker studies like Tasaki et al. regarding drug responses for patients with rheumatoid arthritis, or like Khera et al. on genome-wide polygenic scores as a risk assessment to develop coronary artery disease, type 2 diabetes, atrial fibrillation, inflammatory bowel disease, or breast cancer [67,68].
Gene editing: gene editing biotechnology of clustered regularly interspaced short palindromic repeats (CRISPR) and its associated protein 9 (Cas9) uses short ribonucleic acids (RNAs) as guides (sgRNA) to target a specific deoxyribonucleic acid (DNA) location in order to cut and edit it. These guides, however, may fit DNA locations other than the desired target resulting in the so-called off-target effect. Thus, the selection of the sgRNA molecules to be used is of significant importance. Machine learning algorithms have proved to be promising in the identification of such molecules caring the lowest possible off-target propensity for specific DNA targets [69,70].
Personalized medicine: patients´ symptoms, signs, and test results have to be evaluated by a physician or a multidisciplinary team of experts before a treatment plan is suggested. International Business Machines Corporation (IBM) AI platform “Watson” was initially made known by winning a television quiz show competition. In a study by Wrzeszczynski et al., Watson managed, in 10 minutes, to deliver a treatment plan for a glioblastoma case comparable to the plan that experts made in 160 hours [71]. In another study, Watson was able to suggest cancer therapeutic options that oncologists had overlooked [72]. It seems that if AI systems are provided with large enough amount of data, then they may outperform human physicians in diagnoses or treatment plans. The challenge becomes more intense when big data, such as omics, microbiome sequencing, EHR, social media, and digital images and videos are implemented to the patients' care. Big data are heterogeneous and continuously adding up. As a result, it is difficult for humans to manually analyze them in an effective and meaningful manner in t he field of healthcare. In contrast, AI has the potential to undertake and deliver this task. Interesting approaches are the web-based AI platforms or AI smartphone applications which answer patients´ questions, provide them with advice on whether their condition requires medical attention, and monitor adherence to medications [73,74].
Artificial intelligence research is expanding, and there are increasing AI applications in medicine, too. It is a quickly evolving new era given that DL algorithms seem to perform better than statistics or humans, especially when it comes to big data. Artificial intelligence is a valuable tool, firstly and most importantly, for people and their healthcare. As such, physicians and healthcare systems will embrace, adapt, and evolve accordingly. It is becoming more and more apparent that AI will eventually create the pre- and post- AI era in medicine, too.
The authors declare no competing interests.
Andreas Larentzakis and Nik Lygeros conceived the idea for the manuscript, reviewed the literature, drafted the manuscript, and gave approval of the final version. All the authors have read and agreed to the final manuscript.
Figure 1: deep neural network architecture
Figure 2: single artificial neuron with three inputs and one output
- Turing AM. Computing macherinery and inteligence. Mind. 1950;236:433-460. Google Scholar
- McCulloch WS, Pitts W. A logical calculus of the ideas immanent in nervous activity. Bull Math Biol. 1943;5(4):115-133. PubMed | Google Scholar
- Wiener N. Cybernetics: or control and communication in the animal and the machine. New York and Paris. John Wiley and Sons and Hermann et Cie. 1948.
- Mintz Y, R Brodie. Introduction to artificial intelligence in medicine. Minim Invasive Ther Allied Technol. 2019;28(2):73-81. PubMed | Google Scholar
- Le QV, Ranzato MA, Monga R, Devin M, Chen K, Corrado GS et al. Building high-level features using large scale unsupervised learning. arXiv. 2011;1112:6209.
- Bengio Y, Courville A, Vincent P. Representation learning: a review and new perspectives. IEEE Trans Pattern Anal Mach Intell. 2013;35(8):1798-828. PubMed | Google Scholar
- LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015 May 28;521(7553):436-44. PubMed | Google Scholar
- Schmidhuber J. Deep learning in neural networks: an overview. Neural Netw. 2015 Jan;61:85-117. PubMed | Google Scholar
- Jouppi N. Google supercharges machine learning tasks with TPU custom chip. 2016. accessed 27th March 2020.
- Silver D, Huang A, Maddison CJ, Guez A, Sifre L, van den Driessche G et al. Mastering the game of go with deep neural networks and tree search. Nature. 2016 Jan 28;529(7587):484-9. PubMed | Google Scholar
- Silver D, Hubert T, Schrittwieser J, Antonoglou I, Lai M, Guez A et al. Mastering chess and shogi by self-play with a general reinforcement learning algorithm. Science. 2018 Dec 7;362(6419):1140-1144. PubMed | Google Scholar
- Silver D, Schrittwieser J, Simonyan K, Antonoglou I, Huang A, Guez A et al. Mastering the game of go without human knowledge. Nature. 2017 Oct 18;550(7676):354-359. PubMed | Google Scholar
- Silver D, Hubert T, Schrittwieser J, Antonoglou I, Lai M, Guez A et al. A general reinforcement learning algorithm that masters chess, shogi, and go through self-play. Science. 2018 Dec 7;362(6419):1140-1144 PubMed | Google Scholar
- Senior AW, Evans R, Jumper J, Kirkpatrick J, Sifre L, Green T et al. Protein structure prediction using multiple deep neural networks in the 13th critical assessment of protein structure prediction (CASP13). Proteins: structure, function, and bioinformatics. 2019 Dec;87(12):1141-1148. PubMed | Google Scholar
- Senior AW, Evans R, Jumper J, Kirkpatrick J, Sifre L, Green T et al. Improved protein structure prediction using potentials from deep learning. Nature. 2020 Jan;577(7792):706-710. PubMed | Google Scholar
- Cohen JP, Bertin P, Frappier V. Chester: a web delivered locally computed chest X-ray disease prediction system. arXiv. 2020;1901.11210. Google Scholar
- Lakhani P, Sundaram B. Deep learning at chest radiography: automated classification of pulmonary tuberculosis by using convolutional neural networks. Radiology. 2017 Aug;284(2):574-582. PubMed | Google Scholar
- Halabi SS, Prevedello LM, Kalpathy-Cramer J, Mamonov AB, Bilbily A, Cicero M et al. The radiological society of North America pediatric bone age machine learning challenge. Radiology. 2019 Feb;290(2):498-503. PubMed | Google Scholar
- Thian YL, Li Y, Jagmohan P, Sia D, VEY Chan, Tan RT. Convolutional neural networks for automated fracture detection and localization on wrist radiographs. Radiology: Artificial Intelligence. 2019;1(1):e180001. Google Scholar
- Larson DB, Chen MC, Lungren MP, Halabi SS, Stence NV, Langlotz CP et al. Performance of a deep-learning neural network model in assessing skeletal maturity on pediatric hand radiographs. Radiology. 2018 Apr;287(1):313-322. PubMed | Google Scholar
- Gonzalez G, Ash SY, Vegas-Sanchez-Ferrero G, Onieva JO, Rahaghi FN, Ross JC et al. Disease staging and prognosis in smokers using deep learning in chest computed tomography. Am J Respir Crit Care Med. 2018 Jan 15;197(2):193-203. PubMed | Google Scholar
- Gargeya R, Leng T. Automated identification of diabetic retinopathy using deep learning. Ophthalmology. 2017 Jul;124(7):962-969. PubMed | Google Scholar
- Abramoff MD, Lavin PT, Birch M, Shah N, Folk JC. Pivotal trial of an autonomous AI-based diagnostic system for detection of diabetic retinopathy in primary care offices. NPJ Digit Med. 2018 Aug 28;1:39. PubMed | Google Scholar
- van der Heijden AA, Abramoff MD, Verbraak F, van Hecke MV, Liem A, Nijpels G et al. Validation of automated screening for referable diabetic retinopathy with the IDx-DR device in the horn diabetes care system. Acta Ophthalmol. 2018 Feb;96(1):63-68. PubMed | Google Scholar
- De Fauw J, Ledsam JR, Romera-Paredes B, Nikolov S, Tomasev N, Blackwell S et al. Clinically applicable deep learning for diagnosis and referral in retinal disease. Nat Med. 2018 Sep;24(9):1342-1350 PubMed | Google Scholar
- Long E, Lin H, Liu H, Wu X, Wang L, Jiang J et al. An artificial intelligence platform for the multihospital collaborative management of congenital cataracts. Nature Biomedical Engineering. 2017;1(2):0024. Google Scholar
- Burlina PM, Joshi N, Pekala M, Pacheco KD, Freund DE, Bressler NM et al. Automated grading of age-related macular degeneration from color fundus images using deep convolutional Neural Networks. Jama Ophthalmology. 2017 Nov 1;135(11):1170-1176. PubMed | Google Scholar
- Laukamp KR, Thiele F, Shakirin G, Zopfs D, Faymonville A, Timmer M et al. Fully automated detection and segmentation of meningiomas using deep learning on routine multiparametric MRI. Eur Radiol. 2019 Jan;29(1):124-132.. PubMed | Google Scholar
- Ehteshami Bejnordi B, Veta M, Johannes van Diest P, van Ginneken B, Karssemeijer P, Litjens G et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women With breast cancer. JAMA. 2017 Dec 12;318(22):2199-2210. PubMed | Google Scholar
- Liu Y,Kohlberger T, Norouzi M, Dahl GE, Smith JL, Mohtashamian A et al. Artificial intelligence-based breast cancer nodal metastasis detection: insights into the black box for pathologists. Arch Pathol Lab Med. 2019 Jul;143(7):859-868. PubMed | Google Scholar
- Steiner DF, MacDonald R, Liu Y, Truszkowski P,Hipp JD, Gammage C et al. Impact of deep learning assistance on the histopathologic review of lymph nodes for metastatic breast cancer. Am J Surg Pathol. 2018 Dec;42(12):1636-1646. PubMed | Google Scholar
- McKinney SM, Sieniek M, Godbole V, Godwin J, Antropova N, Ashrafian H et al. International evaluation of an AI system for breast cancer screening. Nature. 2020 Jan;577(7788):89-94. PubMed | Google Scholar
- Soffer S, Ben-Cohen A, Shimon O, Amitai MM, Greenspan H, Klang E et al. Convolutional neural networks for radiologic images: a radiologist's guide. Radiology. 2019 Mar;290(3):590-606. PubMed | Google Scholar
- Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017 Feb 2;542(7639):115-118. PubMed | Google Scholar
- Haenssle HA, Fink C, Schneiderbauer R, Toberer F, Buhl T, Blum A et al. Man against machine: diagnostic performance of a deep learning convolutional neural network for dermoscopic melanoma recognition in comparison to 58 dermatologists. Ann Oncol. 2019 Feb 1;30(2):130e. PubMed | Google Scholar
- Brinker TJ, Hekler A, Enk AH, Klode J, Hauschild A, Berking C et al. Deep learning outperformed 136 of 157 dermatologists in a head-to-head dermoscopic melanoma image classification task. Eur J Cancer. 2019 May;113:47-54. PubMed | Google Scholar
- Brinker TJ, Hekler A, Enk AH, Klode J, Hauschild A, Berking C et al. A convolutional neural network trained with dermoscopic images performed on par with 145 dermatologists in a clinical melanoma image classification task. Eur J Cancer. 2019 Apr;111:148-154. PubMed | Google Scholar
- Brinker TJ, Hekler A, Hauschild A, Berking C, Schilling B, Enk AH et al. Comparing artificial intelligence algorithms to 157 German dermatologists: the melanoma classification benchmark. Eur J Cancer. 2019 Apr;111:30-37. PubMed | Google Scholar
- Hamm CA, Wang CJ, Savic LJ, Ferrante M, Schobert I, Schlachter T et al. Deep learning for liver tumor diagnosis part I: development of a convolutional neural network classifier for multi-phasic MRI. Eur Radiol. 2019 Jul;29(7):3338-3347. PubMed | Google Scholar
- Kawauchi K, Furuya S, Hirata K, Katoh C, Manabe O, Kobayashi K et al. A convolutional neural network-based system to classify patients using FDG PET/CT examinations. BMC Cancer. 2020 Mar 17;20(1):227. PubMed | Google Scholar
- Hirasawa T, Aoyama K, Tanimoto T, Ishihara S, Shichijo S, Ozawa T et al. Application of artificial intelligence using a convolutional neural network for detecting gastric cancer in endoscopic images. Gastric Cancer. 2018 Jul;21(4):653-660. PubMed | Google Scholar
- Horie Y, Yoshio T, Aoyama K, Yoshimizu Y Horiuchi S, Ishiyama A et al. Diagnostic outcomes of esophageal cancer by artificial intelligence using convolutional neural networks. Gastrointestinal Endoscopy. 2019 Jan;89(1):25-32. PubMed | Google Scholar
- Byrne MF, Chapados N, Soudan F, Oertel C, Perez ML, Kelly R et al. Real-time differentiation of adenomatous and hyperplastic diminutive colorectal polyps during analysis of unaltered videos of standard colonoscopy using a deep learning model. Gut. 2019 Jan;68(1):94-100. PubMed | Google Scholar
- Weng SF, Reps J, Kai J,Garibaldi JM, Qureshi N. Can machine-learning improve cardiovascular risk prediction using routine clinical data? .PLoS One. 2017 Apr 4;12(4):e0174944 PubMed | Google Scholar
- Al'Aref SJ,Anchouche K, Singh G, Slomka PJ, Kolli KK, Kumar A et al. Clinical applications of machine learning in cardiovascular disease and its relevance to cardiac imaging. Eur Heart J. 2019 Jun 21;40(24):1975-1986. PubMed | Google Scholar
- Attia ZI, Kapa S, Lopez-Jimenez F, McKie PM, Ladewig DJ, Satam G et al. Screening for cardiac contractile dysfunction using an artificial intelligence-enabled electrocardiogram. Nat Med. 2019 Jan;25(1):70-74. PubMed | Google Scholar
- Dey D, Slomka PJ, Leeson P,Comaniciu D, Shrestha S, Sengupta PP et al. Artificial intelligence in cardiovascular imaging Journal of the American College of Cardiology state-of-the-art review. J Am Coll Cardiol. 2019 Mar 26;73(11):1317-1335. PubMed | Google Scholar
- Dawes TJW, de Marvao A, Shi W, Fletcher T, Watson GMJ, Wharton J et al. Machine Learning of three-dimensional right ventricular motion enables outcome prediction in pulmonary hypertension: a cardiac MR imaging study. Radiology. 2017 May;283(2):381-390. PubMed | Google Scholar
- Arbabshirani MR, Fornwalt BK, Mongelluzzo GJ, Suever JD, Geise BD, Patel AA et al. Advanced machine learning in action: identification of intracranial hemorrhage on computed tomography scans of the head with clinical workflow integration. NPJ Digit Med. 2018 Apr 4;1:9. PubMed | Google Scholar
- Gheiratmand M, Rish I, Cecchi GA, Brown MRG, Greiner R, Polosecki PI et al. Learning stable and predictive network-based patterns of schizophrenia and its clinical symptoms. NPJ Schizophr. 2017 May 16;3:22. PubMed | Google Scholar
- Hazlett HC, H Gu H, Munsell BC, Kim SH, Styner M, Wolff JJ et al. Early brain development in infants at high risk for autism spectrum disorder. Nature. 2017 Feb 15;542(7641):348-351. PubMed | Google Scholar
- Just MA, Pan L, Cherkassky VL, McMakin DL, C Cha C, Nock MK et al. Machine learning of neural representations of suicide and emotion concepts identifies suicidal youth. Nat Hum Behav. 2017;1:911-919. PubMed | Google Scholar
- Mathotaarachchi S, Pascoal TA, Shin M, Benedet AL, Kang MS, Beaudry T et al. Identifying incipient dementia individuals using machine learning and amyloid imaging. Neurobiol Aging. 2017 Nov;59:80-90. PubMed | Google Scholar
- Walsh CG, Ribeiro JD, Franklin JC. Predicting risk of suicide attempts over time through machine learning. Clinical Psychological Science. 2017;5(3):457-469. Google Scholar
- Ho CS, Jean N, Hogan CA, Blackmon L, Jeffrey SS, Holodniy M et al. Rapid identification of pathogenic bacteria using Raman spectroscopy and deep learning. Nat Commun. 2019 Oct 30;10(1):4927 PubMed | Google Scholar
- Mayhew MB, L Buturovic, R Luethy, U Midic, AR Moore, JA Roque et al. A generalizable 29-mRNA neural-network classifier for acute bacterial and viral infections. Nat Commun. 2020 Mar 4;11(1):1177. PubMed | Google Scholar
- Fleck DE, Ernest N, Adler CM, Cohen K, Eliassen JC, Norris M et al. Prediction of lithium response in first-episode mania using the lithium intelligent agent (LITHIA): pilot data and proof-of-concept. Bipolar Disord. 2017 Jun;19(4):259-272. PubMed | Google Scholar
- Lucas GM, Rizzo A, Gratch J, Scherer S, Stratou G, Boberg J et al. Reporting mental health symptoms: breaking down barriers to care with virtual human interviewers. Frontiers in Robotics and AI. 2017;4(51). Google Scholar
- Shademan A, Decker RS, Opfermann JD, Leonard S, Krieger A, Kim PWC et al. Supervised autonomous robotic soft tissue surgery. Sci Transl Med. 2016 May 4;8(337):337ra64. PubMed | Google Scholar
- Yan A. Chinese robot dentist is first to fit implants in patient�s mouth without any human involvement. 2017. Accessed 08th May, 2020.
- Stephenson N, Shane E, Chase J, Rowland J, Ries D, Justice N et al. Survey of machine learning techniques in drug Discovery. Curr Drug Metab. 2019;20(3):185-193. PubMed | Google Scholar
- Schneider G. Automating drug discovery. Nat Rev Drug Discov. 2018 Feb;17(2):97-113. PubMed | Google Scholar
- Merk D, L Friedrich, F Grisoni, G Schneider. De novo design of bioactive small Molecules by rrtificial Intelligence. Mol Inform. 2018 Jan;37(1-2):1700153. PubMed | Google Scholar
- Stokes JM, Yang K, Swanson K, Jin W, Cubillos-Ruiz A, Donghia NM et al. A Deep learning approach to antibiotic discovery. Cell. 2020 Feb 20;180(4):688-702.e13. PubMed | Google Scholar
- Graves J, Byerly J, Priego E, Makkapati N, Parish SV, Medellin B et al. A Review of deep learning methods for antibodies. Antibodies (Basel). 2020 Apr 28;9(2):12. PubMed | Google Scholar
- Boyiadzis MM, Kirkwood JM, Marshall JL, Pritchard CC, Azad NS, Gulley JL et al. Significance and implications of FDA approval of pembrolizumab for biomarker-defined disease. J Immunother Cancer. 2018 May 14;6(1):35. PubMed | Google Scholar
- Khera AV, Chaffin M, Aragam KG, Haas ME, Roselli C, Choi SH et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet. 2018 Sep;50(9):1219-1224. PubMed | Google Scholar
- Tasaki S, Suzuki K, Kassai Y, Takeshita M, Murota A, Kondo Y et al. Multi-omics monitoring of drug response in rheumatoid arthritis in pursuit of molecular remission. Nat Commun. 2018 Jul 16;9(1):2755. PubMed | Google Scholar
- Lin J, Wong KC. Off-target predictions in clustered regularly interspaced short palindromic repeats-Cas9 gene editing using deep learning. Bioinformatics. 2018 Sep 1;34(17):i656-i663. PubMed | Google Scholar
- Liu Q, Cheng X, Liu G, Li B, Liu X. Deep learning improves the ability of sgRNA off-target propensity prediction. BMC Bioinformatics. 2020 Feb 10;21(1):51. PubMed | Google Scholar
- Wrzeszczynski KO, Frank MO, Koyama T, Rhrissorrakrai K, Robine N, Utro F et al. Comparing sequencing assays and human-machine analyses in actionable genomics for glioblastoma. Neurol Genet. 2017 Jul 11;3(4):e164. PubMed | Google Scholar
- Patel NM, Michelini VV, Snell JM, Balu S, Hoyle AP, Parker JS et al. Enhancing next-generation sequencing-guided cancer care through cognitive computing. Oncologist. 2018 Feb;23(2):179-185. PubMed | Google Scholar
- Bartlett J. Boston children's website to feature self-diagnostic tool made by Boston's Buoy. 2018. Accessed on 13th May, 2020.
- Labovitz DL, Shafner L, Reyes Gil M, Virmani D, Hanina A. Using artificial intelligence to reduce the risk of nonadherence in patients on anticoagulation Therapy. Stroke. 2017 May;48(5):1416-1419. PubMed | Google Scholar