A novel variant of genotype 7b hepatitis C virus emphasizing viral hepatitis elimination challenges for sub-Saharan Africa
Mark Wayne Sonderup, Heidi Smuts, Catherine Wendy Spearman
Corresponding author: Mark Wayne Sonderup, Division of Hepatology, Department of Medicine, Faculty of Health Sciences, University of Cape Town and Groote Schuur Hospital, Cape Town, South Africa 
Received: 01 Jul 2020 - Accepted: 20 Jul 2020 - Published: 30 Jul 2020
Domain: Hepatology,Infectious disease
Keywords: Hepatitis C, genotype diversity, elimination programmes
©Mark Wayne Sonderup et al. Pan African Medical Journal (ISSN: 1937-8688). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Mark Wayne Sonderup et al. A novel variant of genotype 7b hepatitis C virus emphasizing viral hepatitis elimination challenges for sub-Saharan Africa. Pan African Medical Journal. 2020;36:232. [doi: 10.11604/pamj.2020.36.232.24726]
Available online at: https://www.panafrican-med-journal.com//content/article/36/232/full
Case report 
A novel variant of genotype 7b hepatitis C virus emphasizing viral hepatitis elimination challenges for sub-Saharan Africa
A novel variant of genotype 7b hepatitis C virus emphasizing viral hepatitis elimination challenges for sub-Saharan Africa
Mark Wayne Sonderup1,&, Heidi Smuts2, Catherine Wendy Spearman1
&Corresponding author
Sub-Saharan Africa has approximately 10.15 million people viraemic with chronic hepatitis C virus infection, extensive genotype and sub-genotype diversity is present, in addition to novel hepatitis C genotypes. Many of the unusual genotypes have extensive baseline resistance associated substitutions with direct acting antiviral therapy treatment outcome data, limited. We report a patient found to have a novel genotype 7b variant with extensive baseline resistance associated substitutions. There is a clear need for a better understanding of the virological characteristics of hepatitis C populations in sub-Saharan Africa to guide best optimal treatment decisions in national hepatitis C elimination programmes.
Sub-Saharan Africa has approximately 14% (˜10.15 million) of the global 71 million people viraemic with chronic hepatitis C virus (HCV) infection [1]. A key feature of HCV in the region is genotype and extensive sub-genotype diversity, with genotype 1 through 5 documented, but regional clustering of certain genotypes occurring e.g. genotype 1 and 4 [2]. Recent data from Uganda and the Democratic Republic of Congo (DRC) has highlighted this sub-genotype diversity as well as finding previously described novel HCV genotypes [3]. An important observation was the wide range of resistance associated substitutions (RAS) observed, especially in novel genotypes. Given limited access to direct acting antiviral (DAA) HCV therapy, treatment outcome data in these patients was not known. In terms of novel HCV genotypes, genotype 7 HCV was first reported in 2015 in four patients from the DRC [4]. It was genetically distinct from genotypes 1 to 6, fulfilled criteria as a new genotype and the prototype sequence (accession number NC_030791/EF108307) was classified as subtype 7a.
In 2016, using next generation sequencing, a genotype 7b subtype was identified in a man from Kinshasa, in the DRC [5]. Similarly a wide range of RAS were detected in both the NS3, NS5A and NS5B of the virus. As treatment outcome data did not exist, the influence of these RAS are unknown. Treatment data for novel HCV genotypes is scarce. In the ASTRAL-1 study evaluating 12 weeks of the combination pan-genotypic DAA therapy sofosbuvir and velpatasvir, a patient originally from the DRC and enrolled as genotype 2a, achieved a sustained virological response (SVR) [6]. During a post-trial analysis, the patient was confirmed as having genotype 7a with phylogenetic analysis demonstrating it to most closely resemble the previously identified genotype 7a patients [7]. Baseline RAS data for this patient is unpublished and not accessible. What is known is that the subtype diversity seen in genotypes 1 and 4 in sub-Saharan Africa does influence SVR rates. This has been observed in recent treatment data that has highlighted reduced responsiveness to standard DAA regimens, notably in genotype 1 (non-1a or 1b subtypes) and in genotype 4, notably subtype 4r [8, 9]. Many of the treatment regimens included the generically available and cost effective NS5A-NS5B DAA combinations of daclatasvir and sofosbuvir or ledipasvir and sofosbuvir.
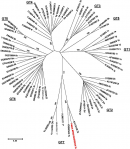
In 2019, a 66-year-old woman from Luanda, Angola was referred to our service. Apart from well controlled hypertension, she was well with no other co-morbidities. Chronic HCV infection was diagnosed 10 years prior, following routine health screening. At that point, genotype analysis performed in Angola reported genotype 2a infection. At that point, she had declined pegylated interferon and ribavirin treatment given its potential toxicities. The route of acquisition of her HCV infection was unknown as no clear identifiable risk factors were evident. Her ALT was 71 IU/L (laboratory normal range 5-40 IU/L) and significant liver fibrosis was excluded with a FibroScanR liver stiffness measurement of 7.6kPa and an AST to Platelet Ratio Index (APRI) score of 0.2. Liver ultrasound was normal and HCV viral load was 9 772 784 IU/mL (Xpert, Cepheid, CA, USA). HIV and HBsAg testing, was negative. We did baseline population-based Sanger sequencing to evaluate for RAS in the NS3, NS5A and NS5B HCV genes using pan genotypic primers and the web-based geno2pheno program (Table 1). Sequences were aligned with GenBank reference sequences using BioEdit and a NS5B neighbour-joining phylogenetic tree was constructed in Mega 6.0 with 1000 bootstrap resamplings (Figure 1). We assigned the sequence as a novel genotype 7b variant (accession number MT633130-MT633134). Considering the presence of extensive baseline RAS in the patient, a DAA therapy regimen was considered (Table 1).
The only available data on treatment was the single genotype 7a patient inadvertently treated in the ASTRAL-1 study [6]. A sofosbuvir/velpatasvir combination was considered, however given the range of RAS, particularly the NS5A 93H mutation, the choice of velpatasvir posed a concern. This mutation impairs treatment response since it induces a high fold in vitro resistance to daclatasvir and velpatasvir. This has been observed to negatively influence SVR in real world experience with DAA therapy, mostly with genotype 3 HCV [10]. An obvious lack of data of the NS5A 93H RAS in genotype 7b in addition to the range of other RAS present, influenced our decision making. Similarly and although not analogous to our patient, data observed in the retreatment MAGELLAN-1 part 2 study suggested that a 16 week duration of treatment with the NS3/NS5A DAA combination of glecaprevir and pibrentasvir, to be better than 12 weeks in patients with prior DAA failure and significant RAS [11]. Given this, we elected to treat the patient with 16 weeks of glecaprevir and pibrentasvir combination as well as the NS5B nucleotide polymerase inhibitor sofosbuvir, all as single daily dosing. After 4 weeks on therapy, HCV viral load was recorded at <10IU/ml (HCV Xpert, Cepheid, CA, USA) and undetected at the end of treatment. Sustained virological response at 12 weeks post end of treatment (SVR12), was confirmed.
We report for the first time an HCV genotype 7b variant. Although this variant is newly described, the patient also underscores challenges in sub-Saharan Africa where a limited, but growing body of experiential treatment data, suggests that prevailing genotype and subtype diversity in countries, may negatively impact SVR rates. This has invariably been observed with standard DAA HCV treatment regimens that notably, are generically available, cost effective and important in building national HCV elimination programmes. Conceding that the DAA regimen used in our patient was possibly overtly aggressive, the patient does represent the dual challenge. Firstly, that there are sub-populations of HCV in sub-Saharan Africa that are less likely to achieve SVR with standard treatment regimens. We have very little knowledge of these unique genotypes and how they are likely to respond to standard therapies. The extensive baseline RAS present in this patient suggests they may be less likely to achieve SVR. Secondly, in our resource-constrained setting, opportunities for re-treatment in the event of non-SVR are problematic and limited by availability, access and cost. Achieving treatment success, the first time around, is crucial. We therefore support the need and would suggest that sub-Saharan African countries sample their HCV population to determine more accurate genotype and sub-genotype characteristics. Such data may serve for better planning in guiding best treatment practice for expanded elimination programmes and thereby potentially limit patients not achieving SVR.
We report a patient with a unique HCV genotype 7b variant. The case provides insight into the extensive HCV genotype diversity in sub-Saharan Africa and highlights the suggested need for countries to gain a better understanding of their HCV populations while developing and implementing national HCV treatment programmes.
The authors declare no competing interests.
MWS authored the report; HS responsible for virological work; CWN provided oversight to manuscript development. All the authors have read and agreed to the final manuscript.
Table 1: baseline NS3, NS5A and NS5B resistance associated substitutions (RAS)
Figure 1: phylogenetic NS5B radial tree comparison of patient (in red) assigned as a novel genotype 7b variant
- Polaris Observatory HCV Collaborators. Global prevalence and genotype distribution of hepatitis C virus infection in 2015: a modelling study. Lancet Gastroenterol Hepatol. 2017;2(3):161-76. PubMed | Google Scholar
- Sonderup MW, Afihene M, Ally R, Apica B, Awuku Y, Cunha L et al. Hepatitis C in sub-Saharan Africa: the current status and recommendations for achieving elimination by 2030. Lancet Gastroenterol Hepatol. 2017;2(12):910-9. PubMed | Google Scholar
- Davis C, Mgomella GS, da Silva Filipe A, Frost EH, Giroux G, Hughes J et al. Highly diverse hepatitis C strains detected in sub-Saharan Africa have unknown susceptibility to direct-acting antiviral treatments. Hepatology. 2019;69(4):1426-41. PubMed | Google Scholar
- Murphy DG, Sablon E, Chamberland J, Fournier E, Dandavino R, Tremblay CL. Hepatitis C virus genotype 7, a new genotype originating from central Africa. J Clin Microbiol. 2015;53(3):967-72. PubMed | Google Scholar
- Salmona M, Caporossi A, Simmonds P, Thélu MA, Fusillier K, Mercier-Delarue S et al. First next-generation sequencing full-genome characterization of a hepatitis C virus genotype 7 divergent subtype. Clin Microbiol Infect. 2016;22(11):947.e1-.e8. PubMed | Google Scholar
- Feld JJ, Jacobson IM, Hézode C, Asselah T, Ruane PJ, Gruener N et al. Sofosbuvir and velpatasvir for HCV genotype 1, 2, 4, 5 and 6 infection. New England Journal of Medicine. 2015;373(27):2599-607. PubMed | Google Scholar
- Schreiber J, McNally J, Chodavarapu K, Svarovskaia E, Moreno C. Treatment of a patient with genotype 7 hepatitis C virus infection with sofosbuvir and velpatasvir. Hepatology. 2016;64(3):983-5. PubMed | Google Scholar
- Childs K, Davis C, Cannon M, Montague S, Filipe A, Tong L et al. Suboptimal SVR rates in African patients with atypical genotype 1 subtypes: implications for global elimination of hepatitis C. J Hepatol. 2019;71(6):1099-105. PubMed | Google Scholar
- Gupta N, Mbituyumuremyi A, Kabahizi J, Ntaganda F, Muvunyi CM, Shumbusho F et al. Treatment of chronic hepatitis C virus infection in Rwanda with ledipasvir-sofosbuvir (SHARED): a single-arm trial. Lancet Gastroenterol Hepatol. 2019;4(2):119-26. PubMed | Google Scholar
- Bertoli A, Sorbo MC, Aragri M, Lenci I, Teti E, Polilli E et al. Prevalence of single and multiple natural NS3, NS5A and NS5B resistance-associated substitutions in hepatitis C virus genotypes 1-4 in Italy. Sci Rep. 2018;8(1):8988. PubMed | Google Scholar
- Poordad F, Pol S, Asatryan A, Buti M, Shaw D, Hézode C et al. Glecaprevir/pibrentasvir in patients with hepatitis C virus genotype 1 or 4 and past direct-acting antiviral treatment failure. Hepatology. 2018;67(4):1253-60. PubMed | Google Scholar